Public Data Access Overview / Example Scripts
We provide I/O example scripts for reading the data files of the Illustris simulations in a few languages. Each has identical functionality, which includes:
- (i) reading a given particle type and/or data field from the snapshot files,
- (ii) reading only the particle subset from the snapshot corresponding to a halo or subhalo,
- (iii) extracting the full subtree or main progenitor branch from either SubLink or LHaloTree for a given subhalo,
- (iv) walking a tree to count the number of mergers,
- (v) reading the entire group catalog at one snapshot,
- (vi) reading specific fields from the group catalog, or the entries for a single halo or subhalo.
We expect they will provide a useful starting point for writing any analysis task, and intend them as a 'minimal working examples' which are short and simple enough that they can be quickly understood and extended.
Currently available are: Python (2.6+ required), IDL (8.0+ required), and Matlab (R2013a+ required). Select one to show all the content on this page specifically for that language.
In all cases, these scripts assume that you have downloaded local copies of the relevant files.
Paths to files are defined with respect to a basePath which is
passed in to all read functions. The locations of group catalog files, snapshot files, and
merger trees files are then specified in a handful of functions, e.g. gcPath(),
snapPath(), and treePath(). These should be modified as necessary to
point to your local files, however you have organized them. By default they assume the following
directory structure: (e.g. for snapshot 68)
- group catalogs:
basePath/groups_068/groups_068.N.hdf5 - snapshots:
basePath/snapdir_068/snap_068.N.hdf5 - SubLink:
basePath/trees/SubLink/tree_extended.N.hdf5 - LHaloTree:
basePath/trees/treedata/trees_sf1_135.N.hdf5
N denotes the file chunk/piece number.
Getting Started Guide
We walk through the process of downloading and exploring data in the Subfind group catalogs, raw snapshot files, and the SubLink merger tree. In these examples we will work with Illustris-3 since the file sizes are smaller to download and easier to work with, although you can replace each occurrence with 'Illustris-1' for the highest resolution run.
If you haven't already, download the
example scripts from Github and make sure they are on your IDL_PATH,
MATLABPATH, or PYTHONPATH, respectively. (For Python, the package can be pip installed).
Group Catalogs
First, make a base directory for the run and a subdirectory for the z=0 group
catalogs (snapshot 135), then download the catalog (~100 MB).
$ mkdir Illustris-3
$ mkdir Illustris-3/groups_135
$ cd Illustris-3/groups_135/
$ wget -nd -nc -nv -e robots=off -l 1 -r -A hdf5 --content-disposition --header="API-Key: INSERT_API_KEY_HERE" "http://www.illustris-project.org/api/Illustris-3/files/groupcat-135/?format=api"
Start up your interface of choice and import the example scripts:
$ python
>>> import illustris_python as il
>>>
$ idl
IDL> .r illustris
% Compiled module: PARTTYPENUM
% Compiled module: (many more)
IDL>
$ matlab
>>
Define the base path for the data (modify as needed), and load the total masses (SubhaloMass) and star formation rate within twice the stellar half mass radius (SubhaloSFRinRad) of all the Subfind subhalos.
>>> basePath = './Illustris-3/'
>>> fields = ['SubhaloMass','SubhaloSFRinRad']
>>> subhalos = il.groupcat.loadSubhalos(basePath,135,fields=fields)
IDL> basePath = "./Illustris-3/"
IDL> fields = ['SubhaloMass','SubhaloSFRinRad']
IDL> subhalos = loadSubhalos(basePath,135,fields=fields)
>> basePath = './Illustris-3/';
>> fields = {'SubhaloMass','SubhaloSFRinRad'};
>> subhalos = illustris.groupcat.loadSubhalos(basePath,135,fields);
Inspecting the return, we see it is a dict/hash/struct with a key count
indicating that there are 121209 total subhalos. Each requested field is returned
as a numeric array with a key name equal to its field name in the group catalog.
>>> subhalos.keys()
['count', 'SubhaloSFRinRad', 'SubhaloMass']
>>> subhalos['SubhaloMass'].shape
(121209,)
IDL> print, subhalos.keys()
count
SubhaloMass
SubhaloSFRinRad
IDL> help, subhalos['SubhaloMass']
<Expression> FLOAT = Array[121209]
>> subhalos
subhalos =
count: 121209
SubhaloMass: [1x121209 single]
SubhaloSFRinRad: [1x121209 single]
Make a simple scatterplot of the relation (note the units).
>>> import matplotlib.pyplot as plt
>>> mass_msun = subhalos['SubhaloMass'] * 1e10 / 0.704
>>> plt.plot(mass_msun,subhalos['SubhaloSFRinRad'],'.')
>>> plt.xscale('log')
>>> plt.yscale('log')
>>> plt.xlabel('Total Mass [$M_\odot$]')
>>> plt.ylabel('Star Formation Rate [$M_\odot / yr$]')
IDL> mass_msun = subhalos['SubhaloMass'] * 1e10 / 0.704
IDL> p = plot(mass_msun, subhalos['SubhaloSFRinRad'], '.', /xlog, /ylog)
IDL> p.xtitle = "Total Mass [$M_{\rm sun}$]"
IDL> p.ytitle = "Star Formation Rate [$M_{\rm sun}$/yr]"
>> mass_msun = subhalos.('SubhaloMass') * 1e10 / 0.704;
>> loglog(mass_msun, subhalos.('SubhaloSFRinRad'), '.');
>> xlabel('Total Mass [$M_\odot$]');
>> ylabel('Star Formation Rate [$M_\odot / yr$]');
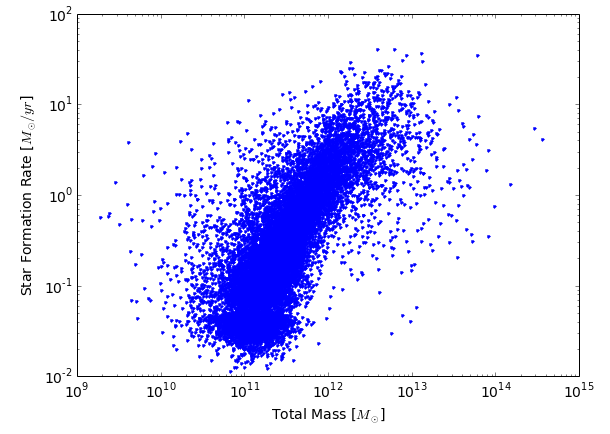
Let us get a list of primary subhalo IDs by loading the GroupFirstSub
field from the FoF groups.
>>> GroupFirstSub = il.groupcat.loadHalos(basePath,135,fields=['GroupFirstSub'])
>>> GroupFirstSub.dtype
dtype('uint32')
>>> GroupFirstSub.shape
(131727,)
IDL> GroupFirstSub = loadHalos(basePath,135,fields=['GroupFirstSub'])
IDL> help, GroupFirstSub
GROUPFIRSTSUB ULONG = Array[131727]
>> GroupFirstSub = illustris.groupcat.loadHalos(basePath,135,{'GroupFirstSub'});
>> class(GroupFirstSub)
ans = uint32
>> size(GroupFirstSub)
ans = 1 131727
Note: Return type
When a single field is requested, the default return is a raw numeric array and not a container type object.For the 5 most massive central subhalos, let's load all their fields from the group catalog and print a gas fraction (gas mass over total baryonic mass) in the stellar half mass radius.
>>> ptNumGas = il.snapshot.partTypeNum('gas') # 0
>>> ptNumStars = il.snapshot.partTypeNum('stars') # 4
>>> for i in range(5):
>>> all_fields = il.groupcat.loadSingle(basePath,135,subhaloID=GroupFirstSub[i])
>>> gas_mass = all_fields['SubhaloMassInHalfRadType'][ptNumGas]
>>> stars_mass = all_fields['SubhaloMassInHalfRadType'][ptNumStars]
>>> frac = gas_mass / (gas_mass + stars_mass)
>>> print GroupFirstSub[i], frac
0 0.0688846
608 0.0236937
1030 0.0638515
1396 0.00357705
1801 0.1222
IDL> ptNumGas = partTypeNum('gas') ; 0
IDL> ptNumStars = partTypeNum('stars') ; 4
IDL> for i=0,4 do begin
IDL> all_fields = loadGroupcatSingle(basePath,135,subhaloID=GroupFirstSub[i])
IDL> gas_mass = all_fields['SubhaloMassInHalfRadType',ptNumGas]
IDL> stars_mass = all_fields['SubhaloMassInHalfRadType',ptNumStars]
IDL> frac = gas_mass / (gas_mass + stars_mass)
IDL> print, GroupFirstSub[i], frac
IDL> endfor
0 0.0688846
608 0.0236937
1030 0.0638515
1396 0.00357705
1801 0.122200
>> ptNumGas = illustris.partTypeNum('gas'); % 0
>> ptNumStars = illustris.partTypeNum('stars'); % 4
>> for i = 1:5
>> all_fields = illustris.groupcat.loadSingle(basePath,135,'subhalo',GroupFirstSub(i));
>> % take care with particle type numbers and 1-based indexing
>> gas_mass = all_fields.('SubhaloMassInHalfRadType')(ptNumGas+1);
>> stars_mass = all_fields.('SubhaloMassInHalfRadType')(ptNumStars+1);
>> frac = gas_mass / (gas_mass + stars_mass);
>> fprintf('%d %f\n', GroupFirstSub(i), frac);
>> end
0 0.068885
608 0.023694
1030 0.063851
1396 0.003577
1801 0.122200
Merger Trees
Let's download the full SubLink merger tree to play with (~8 GB).
$ mkdir Illustris-3/trees
$ mkdir Illustris-3/trees/SubLink
$ cd Illustris-3/trees/SubLink/
$ wget -nd -nc -nv -e robots=off -l 1 -r -A hdf5 --content-disposition --header="API-Key: INSERT_API_KEY_HERE" "http://www.illustris-project.org/api/Illustris-3/files/sublink/?format=api"
Reading individual sub-trees from the full merger trees with 'offsets'
The example scripts make use of pre-computed offsets to accelerate the reading of subsets of the full merger tree files. For example, in order to load the main progenitor branch of the SubLink tree for a single subhalo. These offsets are stored in the group catalog files. Therefore, theloadTree() function requires that you have already downloaded the group catalogs
corresponding to the snapshot of the subhalo (in addition to the tree files themselves).
For the 101st through 105th (indices 100 through 104 if 0-based) most massive primaries, extract the total mass, Subfind ID, and snapshot along the main progenitor branch. Plot the mass histories.
>>> fields = ['SubhaloMass','SubfindID','SnapNum']
>>> start = 100
>>> for i in range(start,start+5):
>>> tree = il.sublink.loadTree(basePath,135,GroupFirstSub[i],fields=fields,onlyMPB=True)
>>> plt.plot(tree['SnapNum'],tree['SubhaloMass'],'-')
>>> plt.yscale('log')
>>> plt.xlabel('Snapshot Number')
>>> plt.ylabel('Total Subhalo Mass [code units]')
IDL> fields = ['SubhaloMass','SubfindID','SnapNum']
IDL> start = 100
IDL> for i=start,start+4 do begin
IDL> tree = loadSublinkTree(basePath,135,GroupFirstSub[i],fields=fields,/onlyMPB)
IDL> p = plot(tree['SnapNum'],tree['SubhaloMass'],'-',/ylog,overplot=(i gt start))
IDL> p.color = (['b','g','r','c','m'])(i-start)
IDL> endfor
IDL> p.xtitle = "Snapshot Number"
IDL> p.ytitle = "Total Subhalo Mass [code units]"
>> fields = {'SubhaloMass','SubfindID','SnapNum'};
>> start = 100;
>> for i = start+1:start+5 % take care with 1-based indexing!
>> tree = illustris.sublink.loadTree(basePath,135,GroupFirstSub(i),fields,true);
>> semilogy(tree.('SnapNum'), tree.('SubhaloMass'), '-');
>> end
>> xlabel('Total Mass [$M_\odot$]');
>> ylabel('Star Formation Rate [$M_\odot$ / yr]');
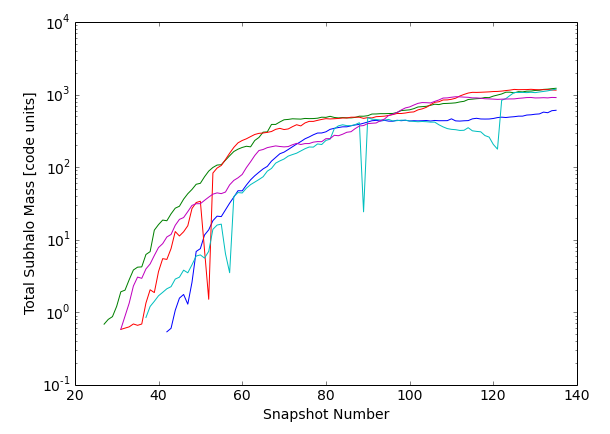
Note that the single-snapshot dips seen in the cyan and red curves can sometimes occur due to the 'subhalo switching problem'. The downward trend in mass followed by the sudden increase in the cyan is a signature of a merger. For details on both aspects of the trees, see the SubLink paper.
We include a semi-complex example of walking through the tree to determine the number of past mergers of a given subhalo, above some mass ratio threshold. Here, the mass ratio is defined as the ratio of the maximum past stellar mass of the two progenitors. For the same halos as above, count the number of major mergers (mass ratio > 1/5).
>>> ratio = 1.0/5.0
>>> # the following fields are required for the walk and the mass ratio analysis
>>> fields = ['SubhaloID','NextProgenitorID','MainLeafProgenitorID','FirstProgenitorID','SubhaloMassType']
>>> for i in range(start,start+5):
>>> tree = il.sublink.loadTree(basePath,135,GroupFirstSub[i],fields=fields)
>>> numMergers = il.sublink.numMergers(tree,minMassRatio=ratio)
>>> print GroupFirstSub[i], numMergers
9106 4
9137 2
9151 3
9170 5
9191 2
IDL> ratio = 1.0/5.0
IDL> ; the following fields are required for the walk and the mass ratio analysis
IDL> fields = ['SubhaloID','NextProgenitorID','MainLeafProgenitorID','FirstProgenitorID','SubhaloMassType']
IDL> for i=start,start+4 do begin
IDL> tree = loadSublinkTree(basePath,135,GroupFirstSub[i],fields=fields)
IDL> numMergers = numMergers(tree,minMassRatio=ratio)
IDL> print, GroupFirstSub[i], numMergers
IDL> endfor
9106 4
9137 2
9151 3
9170 5
9191 2
>> ratio = 1.0/5.0;
>> % the following fields are required for the walk and the mass ratio analysis
>> fields = {'SubhaloID','NextProgenitorID','MainLeafProgenitorID','FirstProgenitorID','SubhaloMassType'};
>> for i = start+1:start+5
>> tree = illustris.sublink.loadTree(basePath,135,GroupFirstSub(i),fields);
>> numMergers = illustris.sublink.numMergers(tree,ratio);
>> fprintf('%d %d\n', GroupFirstSub(i), numMergers);
>> end
9106 4
9137 2
9151 3
9170 5
9191 2
Snapshot Data
Let's download the full z=0 snapshot to play with (~20 GB).
$ mkdir Illustris-3/snapdir_135
$ cd Illustris-3/snapdir_135/
$ wget -nd -nc -nv -e robots=off -l 1 -r -A hdf5 --content-disposition --header="API-Key: INSERT_API_KEY_HERE" "http://www.illustris-project.org/api/Illustris-3/files/snapshot-135/?format=api"
Reading individual halos and subhalos from the snapshots with 'offsets'
As with the merger trees, reading subsets of the snapshots makes use of pre-computed offsets. For example, in order to load only the particles belonging to a single FoF halo. These offsets are stored in the group catalog files. Therefore, theloadSubhalo() and loadHalo() functions require that you have
already downloaded the group catalogs at that snapshot (in addition to the snapshot files
themselves).
First, load the Masses of all the gas cells in the entire box, and calculate
their mean, converting to log solar masses.
>>> import numpy as np
>>> fields = ['Masses']
>>> gas_mass = il.snapshot.loadSubset(basePath,135,'gas',fields=fields)
>>> print np.log10( np.mean(gas_mass,dtype='double')*1e10/0.704 )
7.92160161941
IDL> fields = ['Masses']
IDL> gas_mass = loadSnapSubset(basePath,135,'gas',fields=fields)
IDL> print, alog10( mean(gas_mass)*1e10/0.704 )
7.91676
>> fields = {'Masses'};
>> gas_mass = illustris.snapshot.loadSubset(basePath,135,'gas',fields);
>> gas_mass_mean = sum(gas_mass,'double')/numel(gas_mass); % mean() has significant float error
>> disp( log10( gas_mass_mean*1e10/0.704 ))
7.9216
Next, load the spatial Coordinates of all the dark matter particles in the box,
and make a quick image with a 2D histogram, projecting out the z-axis.
>>> import matplotlib as mpl
>>> dm_pos = il.snapshot.loadSubset(basePath,135,'dm',['Coordinates']);
>>> plt.hist2d(dm_pos[:,0], dm_pos[:,1], norm=mpl.colors.LogNorm(), bins=64);
>>> plt.xlim([0,75000])
>>> plt.ylim([0,75000])
>>> plt.xlabel('x [ckpc/h]')
>>> plt.ylabel('y [ckpc/h]')
IDL> dm_pos = loadSnapshotSubset(basePath,135,'dm',fields=['Coordinates']);
IDL> bin = 75000.0/64
IDL> h = hist_2d(dm_pos[0,*], dm_pos[1,*], bin1=bin,bin2=bin,min1=0,min2=0,max1=75000,max2=75000)
IDL> tv, bytscl(alog10(h>1)), xs=10, ys=10
>> dm_pos = illustris.snapshot.loadSubset(basePath,135,'dm',{'Coordinates'});
>> h = hist3([dm_pos(1,:)' dm_pos(2,:)'], [64 64]);
>> imagesc(log10(h));
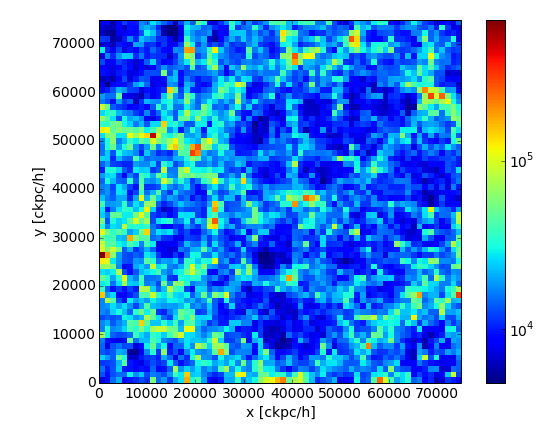
Finally, load the star particles belonging to FoF halo ID 100 (all fields). Print the minimum and maximum of all positions for each axis to check we have loaded only stars in a localized region.
>>> stars = il.snapshot.loadHalo(basePath,135,100,'stars')
>>> stars.keys()
['count', u'GFM_Metals', u'SubfindVelDisp', u'GFM_InitialMass', u'Masses', u'Velocities', u'Coordinates', u'Potential', u'SubfindHsml', u'SubfindDensity', u'NumTracers', u'ParticleIDs', u'GFM_StellarFormationTime', u'GFM_StellarPhotometrics', u'GFM_Metallicity']
>>> for i in range(3):
>>> print np.min(stars['Coordinates'][:,i]), np.max(stars['Coordinates'][:,i])
17993.7 19585.6
58373.0 59606.2
67596.5 68610.6
IDL> stars = loadHalo(basePath,135,100,'stars')
IDL> print, stars.keys()
Masses
GFM_InitialMass
Potential
SubfindHsml
Velocities
Coordinates
count
GFM_StellarFormationTime
GFM_Metallicity
SubfindDensity
GFM_Metals
NumTracers
SubfindVelDisp
GFM_StellarPhotometrics
ParticleIDs
IDL> for i=0,2 do print, minmax( (stars['Coordinates'])[i,*] )
17993.7 19585.6
58373.0 59606.2
67596.5 68610.6
>> stars = illustris.snapshot.loadHalo(basePath,135,100,'stars');
>> stars
stars =
count: 5548
Coordinates: [3x5548 single]
GFM_InitialMass: [1x5548 single]
GFM_Metallicity: [1x5548 single]
GFM_Metals: [9x5548 single]
GFM_StellarFormationTime: [1x5548 single]
GFM_StellarPhotometrics: [8x5548 single]
Masses: [1x5548 single]
NumTracers: [1x5548 uint32]
ParticleIDs: [1x5548 uint64]
Potential: [1x5548 single]
SubfindDensity: [1x5548 single]
SubfindHsml: [1x5548 single]
SubfindVelDisp: [1x5548 single]
Velocities: [3x5548 single]
>> for i = 1:3
>> fprintf('%g %g\n', min(stars.('Coordinates')(i,:)), max(stars.('Coordinates')(i,:)))
>> end
17993.7 19585.6
58373 59606.2
67596.5 68610.6
I/O Scripts Reference
Loading from the FoF and Subfind group catalogs:
The optional fields argument always accepts a string list/array of field names,
which must agree (case-sensitive) to the available datasets in the group catalog. If it is not
specified, all fields will be read and returned, which could be significantly slower.
illustris_python.groupcat.
def loadSubhalos(basePath, snapNum, fields=None):
""" Load all subhalo information from the entire group catalog for one snapshot
(optionally restrict to a subset given by fields). """
def loadHalos(basePath, snapNum, fields=None):
""" Load all halo information from the entire group catalog for one snapshot
(optionally restrict to a subset given by fields). """
def loadHeader(basePath, snapNum):
""" Load the group catalog header. """
def load(basePath, snapNum):
""" Load complete group catalog all at once. """
def loadSingle(basePath, snapNum, haloID=-1, subhaloID=-1):
""" Return complete group catalog information for one halo or subhalo. """
function loadSubhalos(basePath, snapNum, fields=fields)
; Load all subhalo information from the entire group catalog for one snapshot
; (optionally restrict to a subset given by fields).
function loadHalos(basePath, snapNum, fields=fields)
; Load all halo information from the entire group catalog for one snapshot
; (optionally restrict to a subset given by fields).
function loadHeader(basePath, snapNum, chunkNum=chunkNum)
; Load the group catalog header (chunkNum=0 if not specified).
function loadGroupcat(basePath, snapNum)
; Load complete group catalog all at once.
function loadGroupcatSingle(basePath, snapNum, haloID=hID, subhaloID=shID)
; Return complete group catalog information for one halo or subhalo.
illustris.groupcat.
function [result] = loadSubhalos(basePath, snapNum, fields)
% LOADSUBHALOS Load all subhalo information from the entire group catalog for one snapshot
% (optionally restrict to a subset given by fields).
function [result] = loadHalos(basePath, snapNum, fields)
% LOADHALOS Load all halo information from the entire group catalog for one snapshot
% (optionally restrict to a subset given by fields).
function [header] = loadHeader(basePath, snapNum, chunkNum)
% LOADHEADER Load the group catalog header (chunkNum=0 if not specified).
function [gc] = load(basePath, snapNum)
% LOAD Load complete group catalog all at once.
function [result] = loadSingle(basePath, snapNum, type, id)
% LOADSINGLE Return complete group catalog information for one halo or subhalo.
% Type should be one of {'halo','group','subhalo','subgroup'}.
Loading particle-level data from the snapshots:
The optional fields argument always accepts a string list/array of field names,
which must agree (case-sensitive) to the available datasets for that particle type in the
snapshot. If it is not specified, all fields will be read and returned, which could be significantly slower.
The partType argument may either be the particle type number, or one of the recognized
string names, e.g. 'gas', 'stars', 'bhs', or 'dm'.
illustris_python.snapshot.
def loadSubset(basePath, snapNum, partType, fields=None):
""" Load a subset of fields for all particles/cells of a given partType. """
def loadSubhalo(basePath, snapNum, id, partType, fields=None):
""" Load all particles/cells of one type for a specific subhalo
(optionally restricted to a subset fields). """
def loadHalo(basePath, snapNum, id, partType, fields=None):
""" Load all particles/cells of one type for a specific halo
(optionally restricted to a subset fields). """
function loadSnapSubset(basePath, snapNum, partType, fields=fields)
; Load a subset of fields for all particles/cells of a given partType.
function loadSubhalo(basePath, snapNum, id, partType, fields=fields)
; Load all particles/cells of one type for a specific subhalo
; (optionally restricted to a subset fields).
function loadHalo(basePath, snapNum, id, partType, fields=fields)
; Load all particles/cells of one type for a specific halo
; (optionally restricted to a subset fields).
illustris.snapshot.
function [result] = loadSubset(basePath, snapNum, partType, fields)
% LOADSUBET Load a subset of fields for all particles/cells of a given partType.
function [result] = loadSubhalo(basePath, snapNum, id, partType, fields)
% LOADSUBHALO Load all particles/cells of one type for a specific subhalo
% (optionally restricted to a subset fields).
function [result] = loadHalo(basePath, snapNum, id, partType, fields)
% LOADHALO Load all particles/cells of one type for a specific halo
% (optionally restricted to a subset fields).
Loading from the SubLink merger trees:
The optional fields argument always accepts a string list/array of field names,
which must agree (case-sensitive) to the available datasets in the SubLink trees.
If it is not specified, all fields will be read and returned, which could be significantly slower.
If onlyMPB = True, then only the main progenitor branch will be loaded (that is,
only FirstProgenitor links will be followed).
illustris_python.sublink.
def loadTree(basePath, snapNum, id, fields=None, onlyMPB=False):
""" Load portion of Sublink tree, for a given subhalo, in its existing flat format.
(optionally restricted to a subset fields). """
def numMergers(tree, minMassRatio=1e-10, massPartType='stars', index=0):
""" Calculate the number of mergers along the main-progenitor branch, in this sub-tree (optionally above some mass ratio threshold). """
function loadSublinkTree(basePath, snapNum, id, fields=fields, onlyMPB=onlyMPB)
; Load portion of Sublink tree, for a given subhalo, in its existing flat format.
; (optionally restricted to a subset fields).
function numMergers(tree, minMassRatio=1e-10, massPartType='stars', index=0)
; Calculate the number of mergers along the main-progenitor branch, in this sub-tree (optionally above some mass ratio threshold).
illustris.sublink.
function [result] = loadTree(basePath, snapNum, id, fields, onlyMPB, onlyMDB)
% LOADTREE Load portion of Sublink tree, for a given subhalo, in its existing flat format.
% (optionally restricted to a subset fields).
function [numMergers] = numMergers(tree, minMassRatio, massPartType, index)
% NUMMERGERS Calculate the number of mergers, along the main-progenitor branch, in this sub-tree
% (optionally above some mass ratio threshold).
Loading from the LHaloTree merger trees:
The optional fields argument always accepts a string list/array of field names,
which must agree (case-sensitive) to the available datasets in the LHaloTree trees.
If it is not specified, all fields will be read and returned, which could be significantly slower.
If onlyMPB = True, then only the main progenitor branch will be loaded (that is,
only FirstProgenitor links will be followed). On the other hand, if onlyMDB = True, then
only the main descendant branch (towards z=0) will be loaded.
illustris_python.lhalotree.
def loadTree(basePath, snapNum, id, fields=None, onlyMPB=False):
""" Load portion of LHaloTree, for a given subhalo, re-arranging into a flat format. """
function loadLHaloTree(basePath, snapNum, id, fields=fields, onlyMPB=onlyMPB)
; Load portion of LHaloTree, for a given subhalo, re-arranging into a flat format.
illustris.lhalotree.
function [result] = loadTree(basePath,snapNum,id,fields,onlyMPB)
% LOADTREE Load portion of LHaloTree tree, for a given subhalo, re-arranging into a flat format.
General utility functions:
illustris_python.util.
def partTypeNum(partType):
""" Mapping between common names and numeric particle types. """
function partTypeNum(PT)
; Mapping between common names and numeric particle types.
illustris.
function [s] = partTypeNum(pt)
% PARTTYPENUM Mapping between common names and numeric particle types.