Error 121: Unable to synchronously open file (unable to lock file) (Lab)
Arjun Chawla
26 Feb
Hi, i have been loading the Group positions of halos , pretty much the same way for 2 years but i recently ran into a problem which i verified is not due to any updates on my laptop or the network i am on.
So when i use the line :
GroupPos = il.groupcat.loadHalos(basePath,99,fields=['GroupPos'])[3]*aa/hh
i get error : 'OSError: [Errno 121] Unable to synchronously open file (unable to lock file, errno = 121, error message = 'Remote I/O error')'
I also attach a full image of the error.
One of the weird things is i can brute force it. It occurs 9 times out of 10, but 1 time when i run the cell, it goes through and the error doesn't appear.
Since this line is part of functions i create, it gets frustrating as they can fill up the memory as well.
Help with this will be really appreciated.
Thanks
Arjun
Dylan Nelson
26 Feb
This is some sort of problem with a recent change of the server.
(You are running in the Lab, correct?)
There was a recent change to try to fix this (on Feb 26 at roughly 10am EST). Can you try running similar things more now, and let me know if there is any improvement?
Arjun Chawla
27 Feb
Hi Dylan,
Sorry for the late response, things got a bit busy for me last evening. I am running on the Lab and the problem still seems to persist, but thanks for the clarification on why it might be happening.
Arjun
Dylan Nelson
27 Feb
Thanks, I have made one more change.
Can you keep trying now, and let me know if you see any difference?
Arjun Chawla
27 Feb
Hi Dylan, its still the same issue. I tried logging out and logging in as well and close and reopen notebooks, but it didn't work.
Arjun Chawla
27 Feb
Its actually gotten worse. As i mentioned earlier i would run the cell multiple times, and it would work sometimes, now it always gives the error
I ran into the same issue, but I still couldn't resolve it even after setting ' os.environ["HDF5_USE_FILE_LOCKING"]="FALSE" ' in my python code.
In my case, I’m running a standalone .py script from the terminal.
The script contains several functions and an execution block (if __name__ == "__main__": ...).
I set os.environ["HDF5_USE_FILE_LOCKING"] = "FALSE" inside that execution block, but I still encounter the same error during runtime and the script halts.
Do you happen to know what else I should try, or where this environment variable needs to be set for it to take effect?
By the way, terminal is opened in the Lab workspace.
In addition, I move the code into .ipynb but still same problem occurs.
Sincerely,
Sanghyeon
here is error:
Traceback (most recent call last):
File "/home/tnguser/snap_2d_v3.py", line 463, in <module>
snap_with_rvir(row, partType="star", before=False, colorbar=True, scale="proper")
File "/home/tnguser/snap_2d_v3.py", line 380, in snap_with_rvir
subhalo = il.snapshot.loadSubhalo(basePath, snapNum, gid_, partType, fields=["Coordinates", "ParticleIDs"])
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/home/tnguser/illustris_python/snapshot.py", line 191, in loadSubhalo
subset = getSnapOffsets(basePath, snapNum, id, "Subhalo")
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/home/tnguser/illustris_python/snapshot.py", line 158, in getSnapOffsets
with h5py.File(offsetPath(basePath, snapNum), 'r') as f:
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/conda/lib/python3.11/site-packages/h5py/_hl/files.py", line 567, in __init__
fid = make_fid(name, mode, userblock_size, fapl, fcpl, swmr=swmr)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/conda/lib/python3.11/site-packages/h5py/_hl/files.py", line 231, in make_fid
fid = h5f.open(name, flags, fapl=fapl)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "h5py/_objects.pyx", line 54, in h5py._objects.with_phil.wrapper
File "h5py/_objects.pyx", line 55, in h5py._objects.with_phil.wrapper
File "h5py/h5f.pyx", line 106, in h5py.h5f.open
OSError: [Errno 121] Unable to synchronously open file (unable to lock file, errno = 121, error message = 'Remote I/O error')
Sure, here 's a minimal example that reproduces the issue:
import os
import h5py
import illustris_python as il
os.environ["HDF5_USE_FILE_LOCKING"]="FALSE"
basePath = "/home/tnguser/sims.TNG/TNG100-1/output/"
subhalos = il.groupcat.loadSubhalos(basePath, 99, fields=["SubhaloFlag", "SubhaloGrNr"])
subhaloID = np.where(subhalos["SubhaloFlag"])[0]
HosthaloID = subhalos["SubhaloGrNr"][subhaloID]
for subID in subhaloID:
subhalo = il.snapshot.loadSubhalo(basePath, 99, subID, 'star', fields=['ParticleIDs', 'Coordinates'])
...
...
i = N # any number smaller than len(HostHaloID)
halo = il.groupcat.loadSingle(basePath, 99, haloID=HosthaloID[i])
...
As you can see, the script iteratively loads particle-level subhalo data inside a for loop and reads the catalog-level data for a single halo.
The error does not occur deterministically—it can happen at different iterations and not always at the same line.
The common pattern is that it occurs while reading an HDF5 file via h5py (through illustris_python).
I also encountered another related error (not from above example, but similar code configuration):
I note that I didn't touch anything in snapshot.py
Dylan Nelson
18h
Perhaps this environment variable cannot be set like this, if "Environment variable parsed at library startup" (seems to depend on hdf5 version).
Can you instead try to run export HDF5_USE_FILE_LOCKING=FALSE at the terminal, and then run your script in this terminal?
Sanghyeon Han
18h
Dear Nelson,
Thank you for your advice. I will give it a try.
Running my script in the terminal seems to work better, but I’m wondering about the Jupyter Notebook (.ipynb) environment.
As I mentioned before, the error still occurs stochastically, even when I set os.environ["HDF5_USE_FILE_LOCKING"] = "FALSE"
at the top of the notebook (before running any calculation cells).
Hi, i have been loading the Group positions of halos , pretty much the same way for 2 years but i recently ran into a problem which i verified is not due to any updates on my laptop or the network i am on.
So when i use the line :
GroupPos = il.groupcat.loadHalos(basePath,99,fields=['GroupPos'])[3]*aa/hh
i get error : 'OSError: [Errno 121] Unable to synchronously open file (unable to lock file, errno = 121, error message = 'Remote I/O error')'
I also attach a full image of the error.
One of the weird things is i can brute force it. It occurs 9 times out of 10, but 1 time when i run the cell, it goes through and the error doesn't appear.
Since this line is part of functions i create, it gets frustrating as they can fill up the memory as well.
Help with this will be really appreciated.
Thanks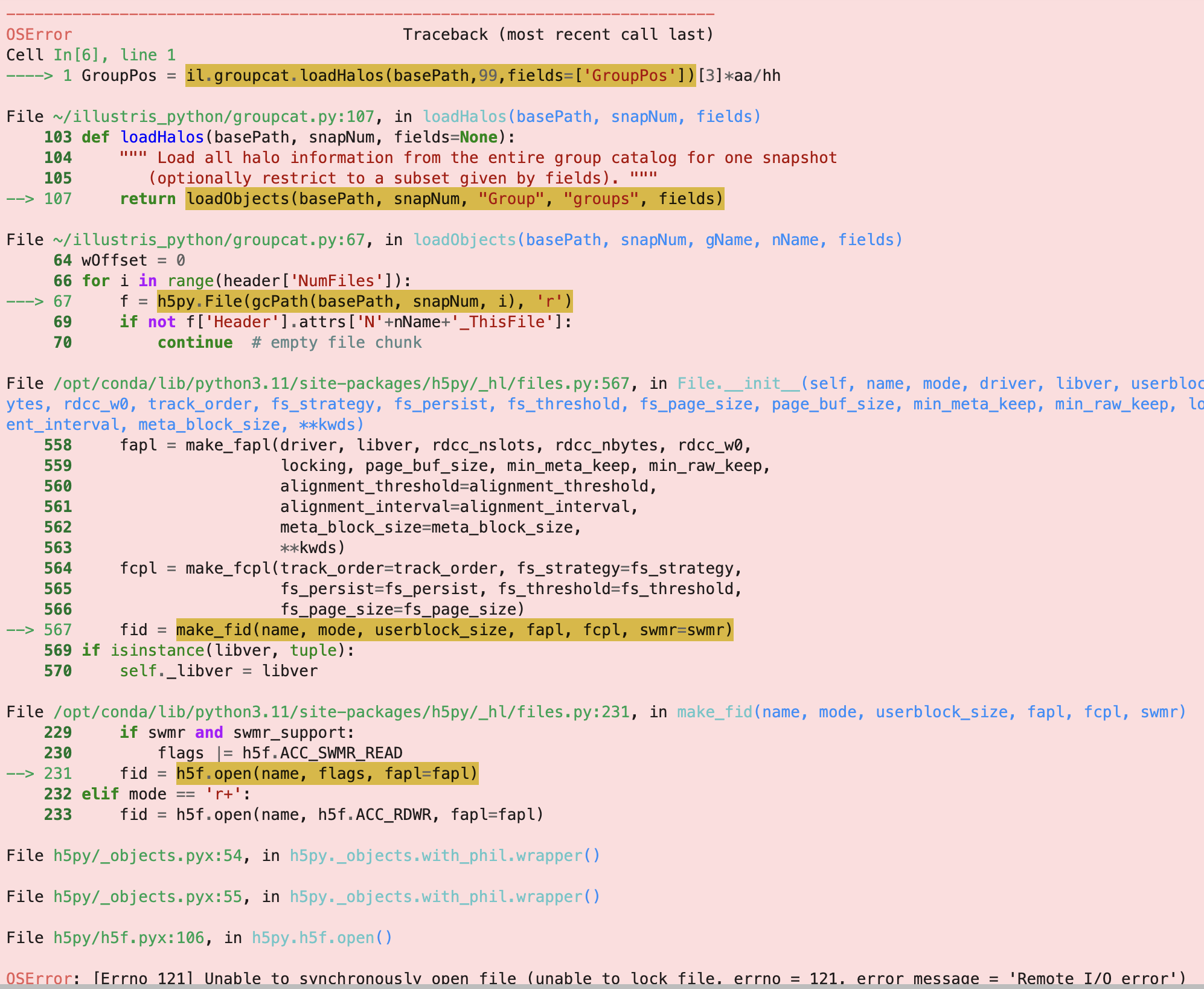
Arjun
This is some sort of problem with a recent change of the server.
(You are running in the Lab, correct?)
There was a recent change to try to fix this (on Feb 26 at roughly 10am EST). Can you try running similar things more now, and let me know if there is any improvement?
Hi Dylan,
Sorry for the late response, things got a bit busy for me last evening. I am running on the Lab and the problem still seems to persist, but thanks for the clarification on why it might be happening.
Arjun
Thanks, I have made one more change.
Can you keep trying now, and let me know if you see any difference?
Hi Dylan, its still the same issue. I tried logging out and logging in as well and close and reopen notebooks, but it didn't work.
Its actually gotten worse. As i mentioned earlier i would run the cell multiple times, and it would work sometimes, now it always gives the error
The issue seems to be related to this:
https://github.com/h5py/h5py/issues/1679
can you try the fix proposed at the bottom?
Hi Dylan, due to me travelling at the moment, i will be able to check and get back to you on Wednesday. Thanks for getting back to me.
Hi Dylan, the fix seems to be working fine. Thanks alot.
Arjun
Thanks for checking, it would be useful if you could please try without this fix, i.e. verify that you still see problems at all?
(I cannot see any similar problems anymore, ever).
Hi Dylan, now i don't see the problem without the fix as well. So its just magically gone .
Dear Nelson,
I ran into the same issue, but I still couldn't resolve it even after setting ' os.environ["HDF5_USE_FILE_LOCKING"]="FALSE" ' in my python code.
In my case, I’m running a standalone .py script from the terminal.
The script contains several functions and an execution block (
if __name__ == "__main__": ...).I set
os.environ["HDF5_USE_FILE_LOCKING"] = "FALSE"inside that execution block, but I still encounter the same error during runtime and the script halts.Do you happen to know what else I should try, or where this environment variable needs to be set for it to take effect?
By the way, terminal is opened in the Lab workspace.
In addition, I move the code into .ipynb but still same problem occurs.
Sincerely,
Sanghyeon
here is error:
Are you still seeing this problem? If so, can you please make a minimal reproducible example?
Dear Nelson,
Sure, here 's a minimal example that reproduces the issue:
As you can see, the script iteratively loads particle-level subhalo data inside a
forloop and reads the catalog-level data for a single halo.The error does not occur deterministically—it can happen at different iterations and not always at the same line.
The common pattern is that it occurs while reading an HDF5 file via
h5py(throughillustris_python).I also encountered another related error (not from above example, but similar code configuration):
I note that I didn't touch anything in
snapshot.pyPerhaps this environment variable cannot be set like this, if "Environment variable parsed at library startup" (seems to depend on hdf5 version).
Can you instead try to run
export HDF5_USE_FILE_LOCKING=FALSEat the terminal, and then run your script in this terminal?Dear Nelson,
Thank you for your advice. I will give it a try.
Running my script in the terminal seems to work better, but I’m wondering about the Jupyter Notebook (.ipynb) environment.
As I mentioned before, the error still occurs stochastically, even when I set
os.environ["HDF5_USE_FILE_LOCKING"] = "FALSE"at the top of the notebook (before running any calculation cells).